Seurat Spatial Transcriptomics. The poly a targeted whole transcriptomics spatial methods are continuously improving the boundaries of cellular resolution. The name of the project which is stored in the seurat object.
For this example we use 10x genomics visium platform brain data. This is the backbone function which takes in single cell expression data to deconvolute spatial transcriptomics spots. In this study, we established human pdac engrafts in mice ischemic hind limbs and used st tissue to investigate st changes in pdac in a hypoxic microenvironment.
It Requires A Prepocessed #' Seurat Object Or A Singlecellexperiment Object As Well As A Rasterised Image #' Saved As An.rds Object.
Researchers can enter this output directly into seurat for further analysis. The bioturing browser is a game changer in the immune oncology field to speed up single cell data sharing among biologists, immunologists and bioinformaticians. Åsa björklund & paulo czarnewski date:
Spaniel Allows Qc Metrics To Be Viewed On Top Of The Histological Image So That Any Quality Issues Can Be Pinpointed.
Assessing the number of genes and number of counts per spot is a useful quality control step. Spatial transcriptomics allows researchers to investigate how gene expression trends varies in space, thus identifying spatial patterns of gene expression. This is a collection of command line r scripts for analyzing spatial transcriptomics data.
New Functionalities Have Been Added That Can Overlay The Transcriptomics Data On The Tissue Histology Image And Provide The Ability To Make An Interactive Plot.
If the object is of class seurat, character string specifying the slot from which to extract the expression matrix. Various methods have been developed for detecting sv (spatially variable) genes, with distinct spatial expression patterns. Saving a seurat object to an h5seurat file is a fairly painless process.
You Can Learn More About The Seurat Workflow For Spatial Gene Expression Data Here →
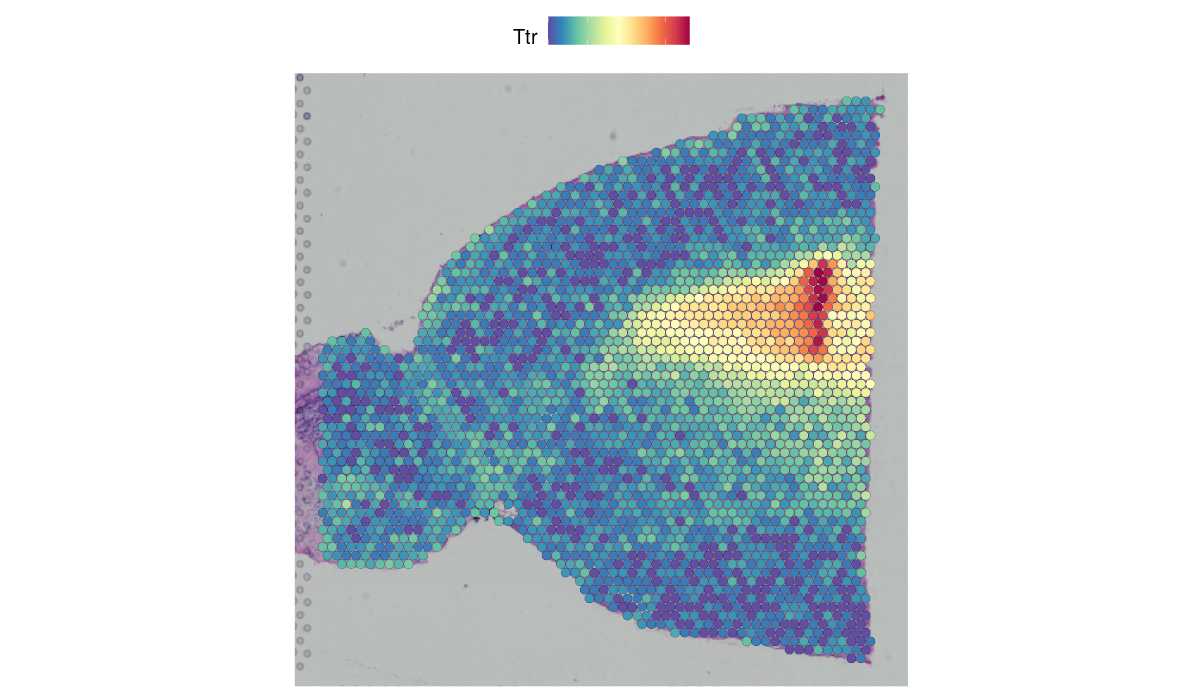
For example, in this data set of the mouse brain, the gene hpca is a strong hippocampus marker and ttr is a marker of the choroid plexus. In this vignette we demonstrate how to build a user friendly workflow starting from data preprocessing, through running mistyr , to analysis of results, i.e., the spatial interactions between markers stored in alternative assays and specific locations. Other topics include the explanation of a general seurat object, and the conversion of sequencing data formats between r and python.
Spatial Transcriptomics Is A Recent Technological Innovation That Measures Transcriptomic Information While Preserving Spatial.
Here we will describe how to use spotlight and seurat label transferring to visualize in stlearn Spatial transcriptomics (st) is a method [1,2,3] that can be used to profile the transcriptome of tissues spatially and is today widely accessible using the 10x genomics visium platform. However, the accuracy of using these sv genes in clustering has not been thoroughly studied.